TreeView Alternatives
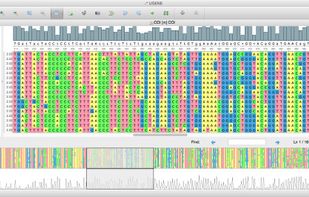
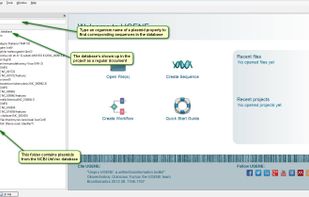
TreeView is described as 'Tree drawing software for Apple Macintosh and Windows' and is an app. There are six alternatives to TreeView for Mac, Windows, Linux and Java. The best TreeView alternative is UGENE, which is both free and Open Source. Other great apps like TreeView are Archaeopteryx, FigTree, TreeView X and Dendroscope.
Alternatives list
Archaeopteryx is a software tool for the visualization, analysis, and editing of potentially large and highly annotated phylogenetic trees.
Cost / License
- Free
- Open Source (LGPL-2.1)
Platforms
- Mac
- Windows
- Linux
- Java
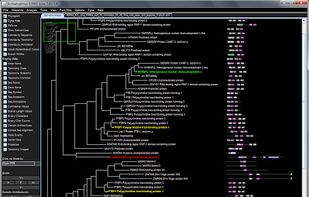
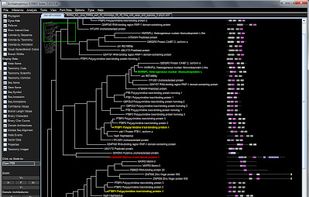
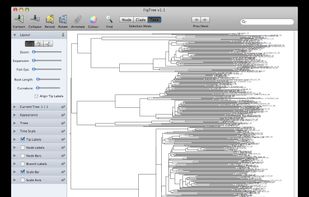
FigTree is designed as a graphical viewer of phylogenetic trees and as a program for producing publication-ready figures. As with most of my programs, it was written for my own needs so may not be as polished and feature-complete as a commercial program.
Cost / License
- Free
- Open Source
Platforms
- Mac
- Windows
- Linux
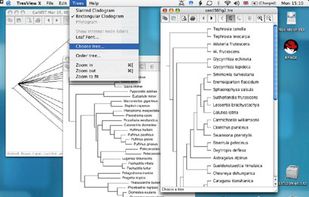
TreeView X is an open source program to display phylogenetic trees on Linux, Unix, Mac OS X, and Windows platforms. It can read and display NEXUS and Newick format tree files (such as those output by PAUP*, .
Cost / License
- Free
- Open Source
Platforms
- Mac
- Windows
- Linux
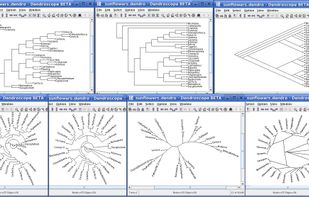
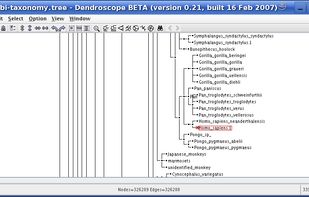
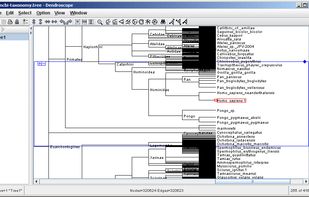
Researchers studying phylogenetic relationships need software that is able to visualize rooted phylogenetic trees and networks efficiently, increasingly of large datasets involving hundreds of thousands of taxa.
Cost / License
- Free
- Proprietary
Platforms
- Mac
- Windows
 +1
+1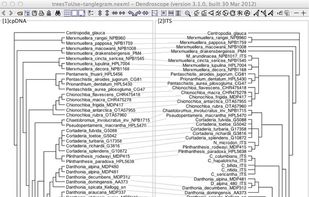
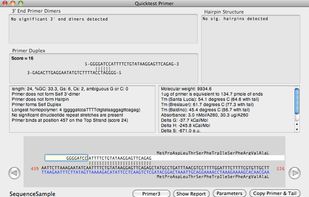
- 6 MacVector alternatives
MacVector is a comprehensive Macintosh application that provides sequence editing, primer design, cloning and designing constructs, internet database searching, protein analysis, sequence confirmation, multiple sequence alignment, phylogenetic reconstruction, coding region...
Cost / License
- Paid
- Proprietary
Platforms
- Mac